From agent-based models to exploring individual microbes and processes that drive species interactions to simulating planetary-scale transports that control ocean biogeography and climate change CBIOMES shared their Julia-based marine microbe and biogeochemical modeling know-how at this year’s Julia programming language conference.
Anchored by CBIOMES’ Gael Forget and Zhen Wu, with additional help from Benoit Pasquier (a postdoc in Seth John’s lab within the Department of Earth Sciences at the University of Southern California, Los Angeles involved in SCOPE) the “Modeling Marine Ecosystems At Multiple Scales Using Julia” session during the recent JuliaCon2021 lifted the lid on a number of new Julia packages of interest to the marine microbe modeling community.
YouTube link to replay the workshop:
https://www.youtube.com/watch?v=UCIRrXz2ZS0
The GitHub repo where you can find & try the notebooks:
https://github.com/JuliaOcean/MarineEcosystemsJuliaCon2021.jl
Packages covered in the workshop included:
- AIBECS.jl: global steady-state biogeochemistry and gridded transport models that run fast for long time scales (centuries or even millennia).
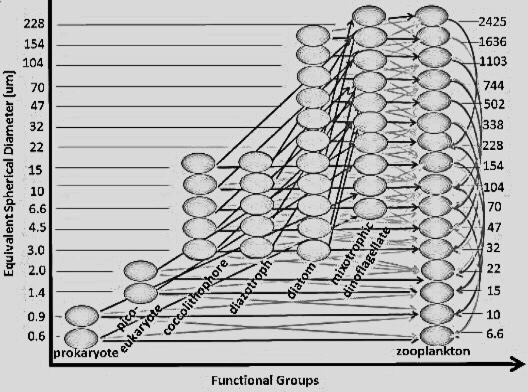
- PlanktonIndividuals.jl: local to global agent-based model, particularly suited to study microbial communities, plankton physiology, and nutrient cycles.
- MITgcmTools.jl: interface to full-featured, Fortran-based, general circulation model and its output (transports, chemistry, ecology, ocean, sea-ice, atmosphere, and more).
- IndividualDisplacements.jl: local to global particle tracking, for simulating dispersion, connectivity, transports in the ocean or atmosphere, etc.
The workshop which was organized around tutorials and self-contained Pluto notebooks for the different packages, followed the following timeline:
- Introduction of the topics covered, presenters, installation, and workshop roadmap (15 minutes).
- AIBECS.jl: concept, implementation, tutorial workthough (30 minutes + 10′ for questions; this notebook URL)
- PlanktonIndividuals.jl: concept, implementation, tutorial workthough (30 minutes + 10′ for questions; this notebook URL)
- ClimateModels.jl and MITgcmToo
ls.jl: concept, implementation, tutorial workthough (20 minutes + 10′ for questions; this notebook URL) - ClimateModels.jl and Individua
lDisplacements.jl: concept, implementation, tutorial workthough (20 minutes + 10′ for questions; this notebook URL) - Q&A, tutorials, etc wrap-up